TRUMPET
TRUMPET: Transcriptional RNA Universal Multi-Purpose gatE plaTform.
The logic gate operations are performed using enzymes acting on DNA template, and the nucleic acig logic readout is fluorescence of RNA aptamers.
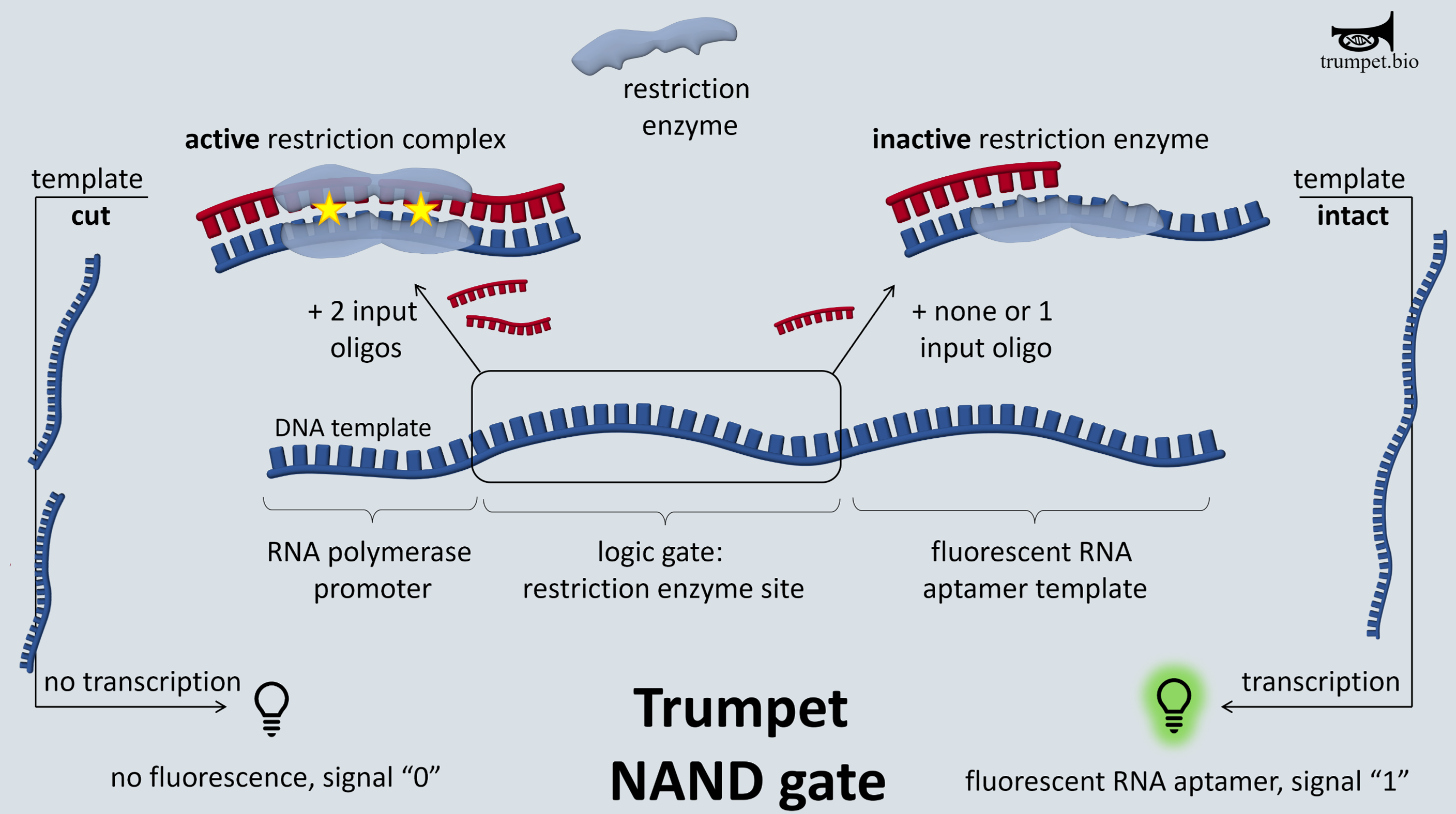
The blueprint of any TRUMPET gate template is a single stranded DNA containing a promoter sequence, a restriction enzyme cut site flanked by random nucleotides, and fluorescent reporter sequence.
The inputs for the gate are short (~20nt) single strands of DNA complementary to one-half of the restriction enzyme cut site and flanking sequence.
By integrating pre-existing secondary structure algorithms — mFold and Nupack — we can robustly verify the compatibility the cut site regions with the downstream RNA fluorescent reporters.
This allows designed gate templates to produce a transcript where the reporter region folds correctly into its secondary structure independent of secondary reactions with the upstream logic gate sequences.
This platform allows to construct complex, layered and interconnected operations on DNA in synthetic cells, with robust fluorescence readout of cell states.
We have used this platform to construct circuits of increased complexity, to investigate the transition from simple biochemical networks to life-like complex genomes.
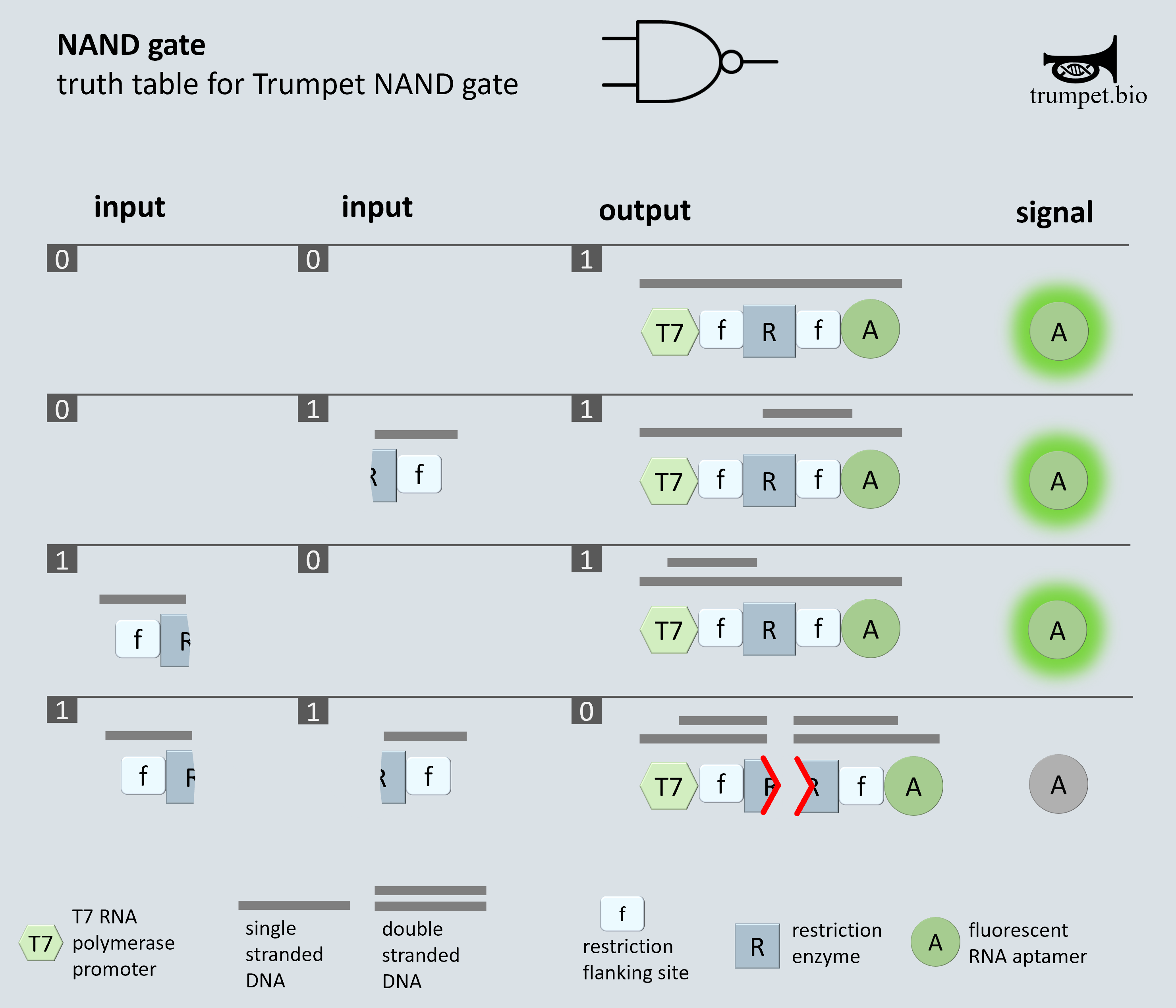
In the use case example of a NAND gate, when both inputs are present in conjunction with a matching gate template, the input sequences anneal to the cut site region on the template, temporarily making the region double stranded. The restriction enzyme cleaves the gate template – preventing the fluorescent reporter sequence from being transcribed. So, with two inputs we have no output signal.
Alternatively, when one or neither inputs are present, the gate template remains intact, and the fluorescent reporter is transcribed. So with one or zero inputs, the output signal is present.
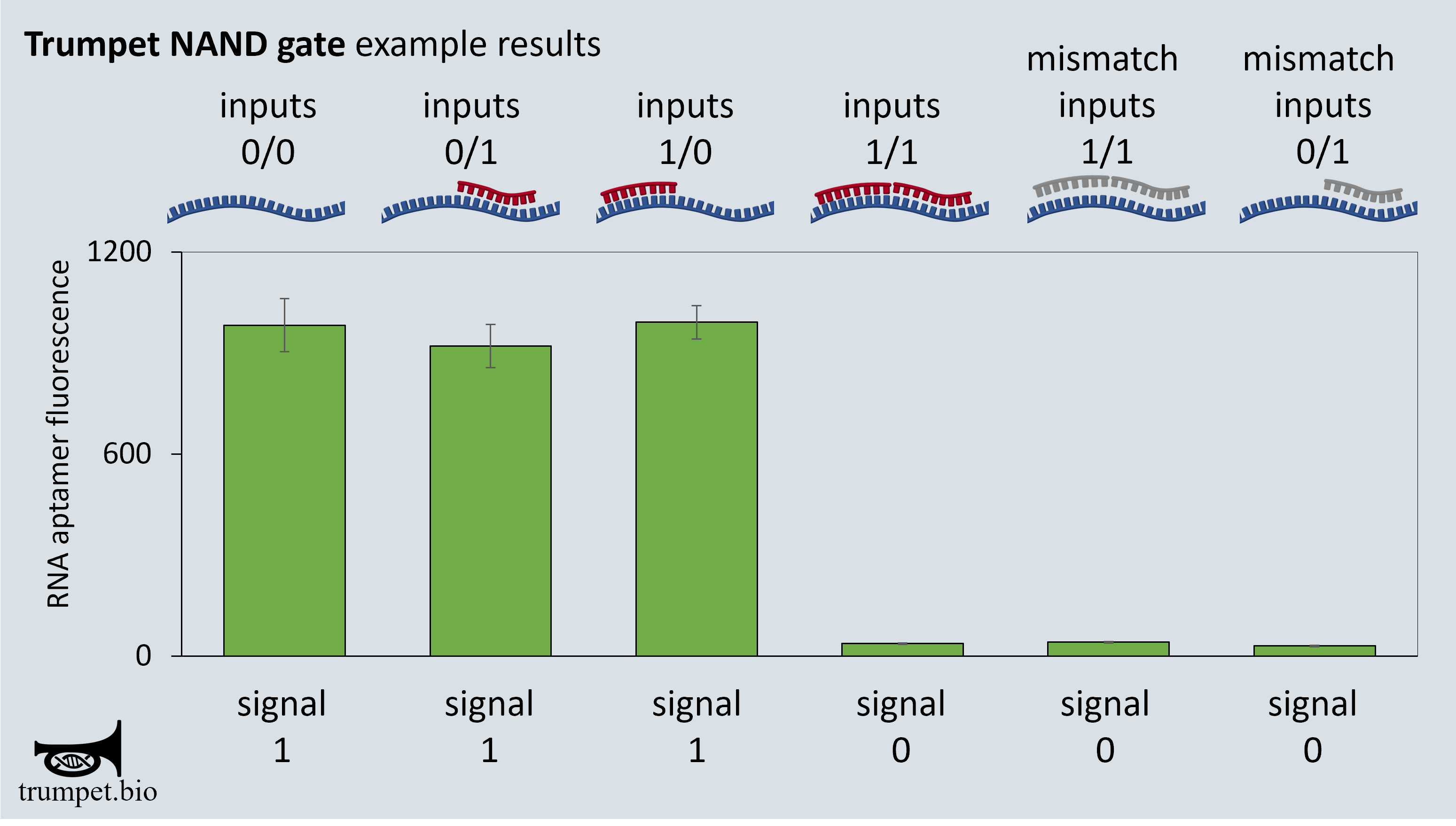
Example data for a NAND gate logic:
Similar operations are possible for all other universal Boolean logic gates within this system.
To cite this work:
Sharon, J.A., Dasrath, C., Fujiwara, A. et al. Trumpet is an operating system for simple and robust cell-free biocomputing. Nat Commun 14, 2257 (2023).
https://doi.org/10.1038/s41467-023-37752-x

Inspiration for the name Trumpet came from Cello, cellocad.org: the first and very well known biological computing platform.
We respect the musical instrument naming convention, and coincidentally two people on team Trumpet play trumpets. The choice was obvious.
Team TRUMPET
The project is part of Adamala lab efforts to engineer toolbox for in vitro biological engineering.
Trumpet team is led by graduate student Judee Sharon, with undergraduate researcher Chelsea Dasrath, and computer scientists Aiden Fujiwara, Alessandro Snyder and Sam O’Brien.
More projects from our lab can be found at protobiology.org.